anti mouse foxm1 (Proteintech)
Structured Review

Anti Mouse Foxm1, supplied by Proteintech, used in various techniques. Bioz Stars score: 96/100, based on 130 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more
https://www.bioz.com/result/anti mouse foxm1/product/Proteintech
Average 96 stars, based on 130 article reviews
Images
1) Product Images from "Abrogation of aberrant glycolytic interactions eliminates senescent cells and alleviates aging-related dysfunctions"
Article Title: Abrogation of aberrant glycolytic interactions eliminates senescent cells and alleviates aging-related dysfunctions
Journal: Signal Transduction and Targeted Therapy
doi: 10.1038/s41392-025-02502-6

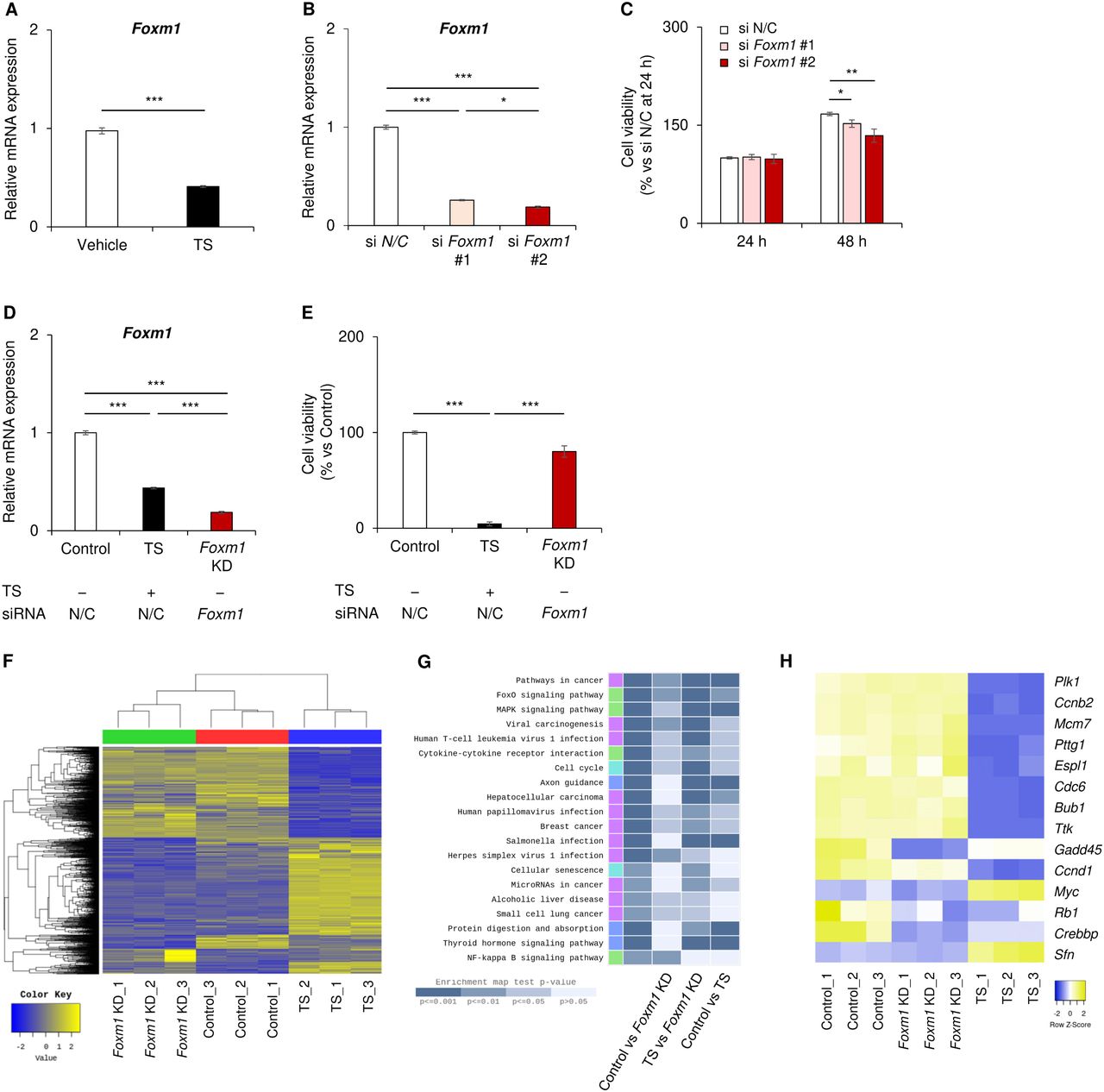
Figure Legend Snippet: A PGAM1-Chk1 antagonist suppresses FOXM1 during senescence. a , b Assessment of the proapoptotic BH family in SnCs after Nutlin 3b treatment. a RT‒PCR analysis of SnCs with or without Nutlin 3b. ( n = 3, biological replicates). b p53 siRNA-transfected SnCs were exposed to Nutlin 3b. RT‒PCR analysis of the indicated cells. c Knockdown of BIM in SnCs. SnCs were transfected with the indicated siRNA, siBIM, or scramble control RNA. The transfectants were then subjected to Nutlin 3b treatment. d – f Comparison of the results of the microarray analysis between the control and Nutlin 3b-treated SnCs ( n = 3, biological replicates). d Fold enrichment analysis by comparison between 1168 genes downregulated by Nutlin 3b in SnCs and 9442 datasets of chromatin immunoprecipitation (ChIP) information reported in the National Center for Biotechnology Information (NCBI), European Bioinformatics Institute (EBI) and DNA Data Bank of Japan (DDBJ). Eighty-six of the 9442 datasets displayed high enrichment (>3-fold). Twenty-one datasets for FOXM1 are shown in red (left panel). e Heatmap of 37 FOXM1 target genes downregulated by Nutlin 3b. SnCs with or without Nutlin 3b were compared. f Gene expression of the FOX family in SnCs after Nutlin 3b treatment, as determined via microarray data. Left panel, heatmap analysis of the FOX family. Right panel: Red and blue bars indicate reduced and increased expression, respectively. Dagger (†) indicates |FC|≥2 and Lpe. P < 0.05 is presented in Supplementary Fig. . Evaluation of FOXM1 protein levels in SnCs after Nutlin 3b treatment ( g ) and after transfection with HIF-2α siRNA ( h ). The data are representative of two independent experiments. i Evaluation of BIM mRNA in SnCs after FOXM1 knockdown. FOXM1 siRNA was transfected into SnCs ( n = 3, biological replicates). j Schematic diagram of the promoter region in the human BIM gene. Several cis-elements for FKH are shown. The indicated regions were subsequently cloned and inserted into luciferase reporter plasmids (left lower panel). Luciferase reporter assays were performed in SnCs after Nutlin 3b treatment (middle panel). A reporter assay for the proximal region was conducted in SnCs after FOXM1 knockdown (right panel) ( n = 3, biological replicates). The data represent the means ± SEMs. Single (*) and double (**) asterisks indicate statistical significance at p < 0.05 and p < 0.01, respectively. Statistical analyses were performed via unpaired Student’s two-tailed t test ( a ) or one-way analysis of variance (ANOVA) and Dunnett’s multiple comparison test ( b , c , i and j )
Techniques Used: Transfection, Knockdown, Control, Comparison, Microarray, Chromatin Immunoprecipitation, Gene Expression, Expressing, Clone Assay, Luciferase, Reporter Assay, Two Tailed Test

Figure Legend Snippet: In vivo FOXM1 accumulation in aged tissues is targeted by Nutlin 3b. a In vivo profiles of FOXM1 mRNA in several tissues of 4-, 10-, 40- and 90-week-old mice, in addition to embryos on day 13.5. The indicated tissues were collected for RNA extraction. b Fold enrichment analysis comparing upregulated genes in 90-week-old mice (liver: 1039; kidney: 1882; lung: 386; and WAT: 385) and ChIP datasets (liver: 9649; kidney: 11,195; lung: 6147; and WAT: 6059). Comparative transcriptomic analysis of the indicated tissues, liver, lung, kidney, and WAT, was performed between 10-week-old and 90-week-old mice. c Heatmap analysis of FOXM1 target genes in the kidney and lung (left and right panels) of young (10-week-old) and aged (90-week-old) mice. d Assessment of FOXM1 target genes involved in the cell cycle and DNA repair genes in aged kidneys and lungs (upper and lower panels, respectively) after Nutlin 3b treatment via RT‒PCR analysis. Tissues were collected from young (10-week-old), aged (90-week-old) and Nutlin 3b-treated aged mice. e The impact of Nutlin-3b on aged kidneys. Representative pictures of the glomerulus after Nutlin 3b treatment (upper panels). Samples from young, aged and Nutlin 3b-treated aged mice. The bar indicates 50 μm. Hyalinosis scores were assessed in aged mice with or without Nutlin 3b treatment. Glomerulus region (lower left) and interstitial region (lower right). f Immunohistochemical examination of the lung for p21 CIP1 . Representative images of stained samples from the indicated mice are shown (left panels). The bar indicates 20 μm. The ratio of p21 CIP1 -positive cells was assessed (right panel). g Comparison of the mRNA levels of p16 Ink4a and SASP factors (Il6, Tnfα, Cxcl1, and Ccl5). Lung extracts from the indicated mice were analyzed via RT‒PCR. The data represent the means ± SEMs. Single (*) and double (**) asterisks indicate statistical significance at p < 0.05 and p < 0.01, respectively. Statistical analyses were performed via one-way analysis of variance (ANOVA) and Dunnett’s multiple comparison test
Techniques Used: In Vivo, RNA Extraction, Immunohistochemical staining, Staining, Comparison